|
Bio-Rad
western blot with anti brca1 ab 1 Western Blot With Anti Brca1 Ab 1, supplied by Bio-Rad, used in various techniques. Bioz Stars score: 85/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/western blot with anti brca1 ab 1/product/Bio-Rad Average 85 stars, based on 1 article reviews
western blot with anti brca1 ab 1 - by Bioz Stars,
2026-03
85/100 stars
|
Buy from Supplier |
|
Bio-Rad
brca1  Brca1, supplied by Bio-Rad, used in various techniques. Bioz Stars score: 91/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/brca1/product/Bio-Rad Average 91 stars, based on 1 article reviews
brca1 - by Bioz Stars,
2026-03
91/100 stars
|
Buy from Supplier |
|
Bio-Rad
monoclonal mouse antibodies  Monoclonal Mouse Antibodies, supplied by Bio-Rad, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/monoclonal mouse antibodies/product/Bio-Rad Average 93 stars, based on 1 article reviews
monoclonal mouse antibodies - by Bioz Stars,
2026-03
93/100 stars
|
Buy from Supplier |
|
Merck & Co
mouse α brca1 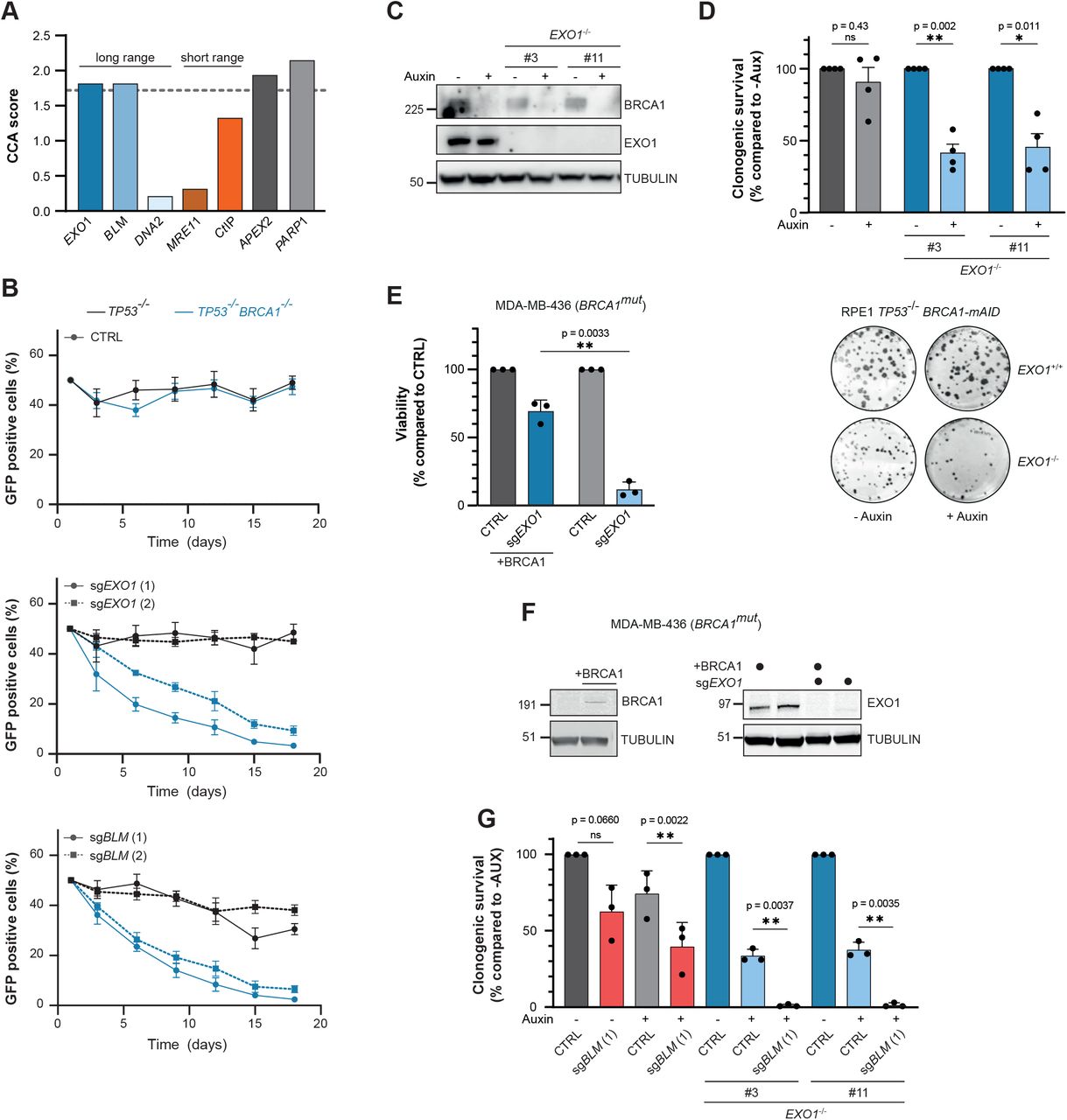 Mouse α Brca1, supplied by Merck & Co, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/mouse α brca1/product/Merck & Co Average 90 stars, based on 1 article reviews
mouse α brca1 - by Bioz Stars,
2026-03
90/100 stars
|
Buy from Supplier |
Image Search Results

Journal: Journal of Cellular and Molecular Medicine
Article Title: Inhibition of c‐MET increases the antitumour activity of PARP inhibitors in gastric cancer models
doi: 10.1111/jcmm.15655
Figure Lengend Snippet: Steady‐state levels of gastric cancer cell lines. Using Western blot assay, steady protein levels of BRCA1, BRCA2 and c‐MET are analysed in primary gastric cancer cell lines HS746T and AGS. Protein levels were normalized against actin
Article Snippet: Antibodies were used against:
Techniques: Western Blot

Journal: Journal of Cellular and Molecular Medicine
Article Title: Inhibition of c‐MET increases the antitumour activity of PARP inhibitors in gastric cancer models
doi: 10.1111/jcmm.15655
Figure Lengend Snippet: Low levels of c‐MET partially sensitize GC cell lines in PARP inhibition. A, HS746T/AGS cells, control‐siRNA‐Hs746T/AGS cells and si‐c‐MET Hs746T/AGS cells were exposed to increasing doses (0‐40 µmol/L) of NU1025 for 48 h for determination of cell viability (MTT metabolic activity assay). The protein levels of c‐MET expression (by Western blot analysis) revealed down‐regulation of the c‐MET receptor in both cell lines (HS746T and AGS); (B) HS746T cells, control‐siRNA‐Hs746T cells and si‐BRCA1/2 Hs746T cells were exposed to increasing doses (0‐40 µmol/L) of NU1025 for 48 h for determination of cell viability (MTT metabolic activity assay). The protein levels of BRCA1 and BRCA2 expression (by Western blot analysis) revealed down‐regulation of the BRCA1/2 in HS746T cell line; (C) HS746T cells, control‐siRNA‐Hs746T, siBRCA1/2‐Hs746T and siMET/BRCA1/2‐Hs746T cells were cultured with the indicated concentrations of NU1025 (5, 10 and 20 μmol/L) for 48 h for determination of cell viability (MTT metabolic activity assay). Error bars represent SD
Article Snippet: Antibodies were used against:
Techniques: Inhibition, Control, Metabolic Assay, Expressing, Western Blot, Cell Culture

Journal: Journal of Cellular and Molecular Medicine
Article Title: Inhibition of c‐MET increases the antitumour activity of PARP inhibitors in gastric cancer models
doi: 10.1111/jcmm.15655
Figure Lengend Snippet: Co‐inhibition of c‐MET (SU11274) and PARP (NU1025) sensitizes GC cells after knockdown BRCA1/2. Knocking down BRCA1 or BRCA2 sensitizes cells to PARP and c‐MET inhibition in HS746T cells expressing low levels of c‐MET (AGS cells, c‐MET knockdown Hs746T cells) to PARP inhibition. A, HS746T cells, control‐siRNA‐Hs746T cells and siBRCA1/2‐Hs746T (upper panel) and AGS (lower panel) cells were exposed to 5 µmol/L of NU1025 and/or 5 µmol/L of SU11274 for 48 h for determination of cell viability (MTT metabolic activity assay). Results are expressed as percentages. Average values of three experiments ± SD are shown; (B) Western blot analysis of PARP and cl.caspase‐3 in Hs746T‐control‐siRNA, siBRCA1/2‐Hs746T (upper panel) and AGS (lower panel) cell lines. Cells were cultured with the indicated drugs (5 μmol/L NU1025, 5 μmol/L SU11274 alone or in combination for 24 h of treatment). Protein levels were normalized against actin
Article Snippet: Antibodies were used against:
Techniques: Inhibition, Knockdown, Expressing, Control, Metabolic Assay, Western Blot, Cell Culture

Journal: Biology of reproduction
Article Title: Dimeric transferrin inhibits phagocytosis of residual bodies by testicular rat Sertoli cells.
doi: 10.1095/biolreprod.107.063107
Figure Lengend Snippet: FIG. 1. Purification and functional char- acterization of dimeric TRF. A) Chromato- graphic profile of highly purified human TRF. The TRF solution was loaded onto an ultragel Aca44 column as described in Materials and Methods. OD, optical density. B) Each fraction was analyzed using native gel electrophoresis and immunoblotting. VLMB, very low-mobility band; LMB, low- mobility band; HMB, high-mobility band. C) Sensorgrams of interaction between high- and low-mobility purified fractions of hTRF and the monoclonal anti-hTRF anti- body. The interaction between both forms of hTRF and the monoclonal antibody is indicated by a dashed line. After reinjection of the monoclonal antibody, only the accessible epitope on the dimeric form generated a secondary signal (dark line), in contrast to the monomeric form (light line). The arrows directed to the top indicate the beginning of injection of the anti-hTRF antibody, and those directed toward the bottom indicate the end of the injections. The principle of discrimination between monomeric versus dimeric TRF by SPR is schematized on the right. RU, resonance units.
Article Snippet:
Techniques: Purification, Functional Assay, Nucleic Acid Electrophoresis, Western Blot, Generated, Injection

Journal: bioRxiv
Article Title: EXO1-mediated DNA repair by single-strand annealing is essential for BRCA1-deficient cells
doi: 10.1101/2023.02.24.529205
Figure Lengend Snippet: (A) Selected results of a gene essentiality screen in BRCA1-proficient and -deficient RPE1 cells ( Adam et al. , 2021 ). Plotted is the CCA score: a higher score indicates a unique essentiality in BRCA1-deficient cells compared to -proficient cells. Dashed line indicates the cut-off for a significant CCA score (based on Adam et al. , 2021 ). (B) RPE1 hTERT cells expressing Cas9, either TP53 -/- (black lines) or TP53 -/- BRCA1 -/- (blue lines), were infected with indicated sgRNA together with GFP, or with an empty vector together with mCherry. GFP- and mCherry-positive cells were mixed 1:1, and the frequency of GFP-positive cells in the population was determined at multiple time points (n=4, mean±SEM). Western blot of lysates shown in supplemental figure 1A. (C) RPE1 hTERT PAC -/- TP53 -/- cells were genetically modified to generate a BRCA1-mAID-GFP fusion gene at the endogenous BRCA1 -locus. In this genetic background, two clonal EXO1 -/- cell lines were generated using gene editing techniques. Cells were treated with auxin (500 μM) for 48 hours or left untreated, and lysates were analysed by western blotting. (D) The cell lines described in were treated with 500 μM auxin, or left untreated, and clonogenic survival was determined. Lower panel shows a representative experiment, top panel shows the quantification (n=4, mean+SEM, *p<0.05, **p<0.01, paired t-test). (E) BRCA1 -mutated MDA-MB-436 cells, either WT or reconstituted with BRCA1, were infected with empty vector (CTRL) or EXO1 -targeting sgRNA and viability was measured using CellTiter-Glo (n=3, mean+SD, **p<0.01, paired t-test). (F) Lysates of the MDA-MB-436 cell lines described in were analysed by western blotting. (G) RPE1 hTERT TP53 -/- BRCA1-mAID-GFP cell lines were virally transduced to express Cas9 cDNA and the indicated gRNAs, followed by a clonogenic survival assay in presence or absence of 500 μM Auxin (n=3, mean+SD, **p<0.01, paired t-test). Western blot of lysates shown in supplemental figure 1L.
Article Snippet: The following primary antibodies were used for western blotting: Mouse α 53BP1 (BD 612522; 1:1,500), Rabbit α BLM (Abcam ab2179; 1:2,000),
Techniques: Expressing, Infection, Plasmid Preparation, Western Blot, Genetically Modified, Generated, Clonogenic Cell Survival Assay

Journal: bioRxiv
Article Title: EXO1-mediated DNA repair by single-strand annealing is essential for BRCA1-deficient cells
doi: 10.1101/2023.02.24.529205
Figure Lengend Snippet: (A) Primary breast tumour samples (Staaf et al. 2019, n=247) were divided in groups based on the presence of biallelic BRCA1 mutations ( BRCA1 mut) or BRCA1 promotor hypermethylation ( BRCA1 prom. MET), and gene expression of indicated genes were plotted. (B) As in , but for another cohort of breast cancer patients in which groups were divided on the presence of deleterious BRCA1 mutations ( Nik-Zainal et al. , 2016 , n=342). (C) For tumour samples of the TCGA breast cancer dataset (n=1,048), the correlation between the HRD score and expression level of an individual gene was determined. Plotted are the Pearson correlation coefficient for each of 60,000 measured transcripts. (D) HRD score and expression of the indicated genes were plotted for each tumour sample in the TCGA breast cancer dataset (n=1,048). Red line is the linear regression curve, PC=Pearson correlation coefficient.
Article Snippet: The following primary antibodies were used for western blotting: Mouse α 53BP1 (BD 612522; 1:1,500), Rabbit α BLM (Abcam ab2179; 1:2,000),
Techniques: Expressing

Journal: bioRxiv
Article Title: EXO1-mediated DNA repair by single-strand annealing is essential for BRCA1-deficient cells
doi: 10.1101/2023.02.24.529205
Figure Lengend Snippet: (A) RPE1 hTERT TP53 -/- BRCA1-mAID-GFP cells, either EXO1 +/+ or EXO1 -/- , were treated with 500 μM auxin for 48 hours to deplete BRCA1 or left untreated. This was followed by Hoechst staining and microscopic quantification of the number of micronuclei (n=5, mean+SD, *p<0.05, ratio paired t-test). (B) RPE1 hTERT TP53 -/- BRCA1 -/- cells expressing Cas9 were infected with sgEXO1 . Next, DAPI-stained nuclei were analysed by microscopy to quantify the number of micronuclei (n=3, mean+SD, ratio paired t-test). (C) RPE1 hTERT TP53 -/- BRCA1-mAID-GFP , either EXO1 +/+ or EXO1 -/- , were treated with 500 μM auxin for 48 hours to deplete BRCA1 or left untreated. Next, metaphase spreads were analysed for chromosomal aberrations. Right panel shows a representative experiment with blue arrows indicating aberrations, left panel shows the quantification (n=3, >40 metaphases per replicate, mean). (D) Data depicted in panel C was re-plotted to show the absolute number of chromosomal aberrations, either chromatid type or chromosome type. (E) RPE1 hTERT TP53 -/- BRCA1-mAID-GFP cells, either EXO1 +/+ or EXO1 -/- , were treated with 500 μM auxin for 48 hours to deplete BRCA1 or left untreated. Nuclear γH2AX intensity in S-phase (EdU + ) cells was analysed by IF microscopy. A representative of two independent experiments is shown, black line indicates median (****p<0.0001, Mann-Whitney). (F) Nuclear γH2AX intensity in S-phase (EdU + ) cells analysed by IF microscopy in BRCA1 -mutated MDA-MB-436 cells, either WT or reconstituted with BRCA1 cDNA, infected with empty vector (CTRL) or EXO1-targeting shRNA. A representative of two independent experiments is shown, black line indicates median (****p<0.0001, Mann-Whitney).
Article Snippet: The following primary antibodies were used for western blotting: Mouse α 53BP1 (BD 612522; 1:1,500), Rabbit α BLM (Abcam ab2179; 1:2,000),
Techniques: Staining, Expressing, Infection, Microscopy, MANN-WHITNEY, Plasmid Preparation, shRNA

Journal: bioRxiv
Article Title: EXO1-mediated DNA repair by single-strand annealing is essential for BRCA1-deficient cells
doi: 10.1101/2023.02.24.529205
Figure Lengend Snippet: (A) RPE1 hTERT TP53 -/- BRCA1 -/- cells expressing Cas9, either TP53BP1 +/+ or TP53BP1 -/- , were transduced with an AAVS1-targeting (CTRL) or EXO1 -targeting sgRNA, followed by a clonogenic survival assay (n=3, mean+SD, **p<0.01, paired t-test). Western blot of lysates shown in supplemental figure 2A. (B) Indicated RPE1 cell lines were incubated with EdU and PARGi for 30 minutes, followed by IF microscopy to analyse PAR formation in EdU-positive cells. A representative of two independent experiments is shown, black line indicates median (****p<0.0001, Mann-Whitney). (C) As in , but for BRCA1 -mutated MDA-MB-436 cells, either WT or reconstituted with BRCA1 cDNA, infected with empty vector (CTRL) or EXO1 -targeting sgRNA. A representative of three independent experiments is shown, black line indicates median (****p<0.0001, Mann-Whitney). (D) As in , but for DLD1 cells, either WT or BRCA2 -/- . A representative of three independent experiments is shown, black line indicates median (****p<0.0001, Mann-Whitney).
Article Snippet: The following primary antibodies were used for western blotting: Mouse α 53BP1 (BD 612522; 1:1,500), Rabbit α BLM (Abcam ab2179; 1:2,000),
Techniques: Expressing, Transduction, Clonogenic Cell Survival Assay, Western Blot, Incubation, Microscopy, MANN-WHITNEY, Infection, Plasmid Preparation

Journal: bioRxiv
Article Title: EXO1-mediated DNA repair by single-strand annealing is essential for BRCA1-deficient cells
doi: 10.1101/2023.02.24.529205
Figure Lengend Snippet: (A) Results of a gene essentiality screen in BRCA1-proficient and -deficient cells ( Adam et al. , 2021 ) were mined to extract the CCA scores for the indicated genes. A higher CCA score indicates a unique essentiality in BRCA1-deficient cells compared to proficient cells. Dashed line indicates the cut-off for a significant CCA score (based on Adam et al. , 2021 ). (B) Indicated RPE1 cells were transfected with a control (siCTRL) or EXO1 -targeting siRNA, followed by treatment with IR and subsequent IF microscopy to analyse either RAD51 or RAD52 foci formation. Left panel shows quantification (n=3, mean+SD, *p<0.05, paired t-test) and right panel shows representative microscopy images of eGFP-RAD52 foci. Western blot of lysates shown in supplemental figure 3A. (C) Clonogenic survival assay of Cas9-expressing RPE1 hTERT TP53 -/- BRCA1 -/- cells that were transduced to express the indicated sgRNAs (n=3, mean+SD, *p<0.05, **p<0.01, one-way ANOVA, post-hoc Dunnett’s, compared to CTRL). Western blot of lysates shown in supplemental figure 3D. (D) HEK 293T cells carrying the DSB-Spectrum_V3 reporter, either WT or EXO1 -/- , were transfected with indicated siRNAs, followed by a second round of transfection with a Cas9 cDNA and BFP sgRNA targeting the reporter locus. Next, cells were analysed by flow cytometry to quantify repair by the indicated pathways (n=4, mean±SEM, *p<0.05, ratio paired t-test). Western blot of lysates shown in supplemental figure 3E. (E) Nuclear γH2AX intensity in S-phase (EdU + ) cells analysed by IF microscopy in BRCA2 -/- cells infected with empty vector or sgEXO1 . A representative of two independent experiments is shown, black line indicates median.
Article Snippet: The following primary antibodies were used for western blotting: Mouse α 53BP1 (BD 612522; 1:1,500), Rabbit α BLM (Abcam ab2179; 1:2,000),
Techniques: Transfection, Microscopy, Western Blot, Clonogenic Cell Survival Assay, Expressing, Flow Cytometry, Infection, Plasmid Preparation

Journal: bioRxiv
Article Title: EXO1-mediated DNA repair by single-strand annealing is essential for BRCA1-deficient cells
doi: 10.1101/2023.02.24.529205
Figure Lengend Snippet: (A) Model of the mechanism causing synthetic lethality between BRCA1-deficiency and EXO1 loss. (B) Whole genome sequencing data of pan cancer tumour samples ( Martínez-Jiménez et al. , 2022 ) was analysed to quantify the number of genetic scars indicative of DSB-repair by SSA, here defined as deletions flanked by homologous sequences of >10bp. CHORD analysis was used to classify samples as HR-proficient or HR-deficient, either BRCA1-type or BRCA2-type ( Nguyen et al. , 2020 ) (****p<0.0001, kolmogornov-smirnov). (C) Tumour samples from a pan-cancer cohort were binned based on SSA scar count, and the EXO1 expression was plotted for each tumour sample. (D) EXO1 expression levels in BRCA1 WT or BRCA1 mutant pan-cancer tumour samples.
Article Snippet: The following primary antibodies were used for western blotting: Mouse α 53BP1 (BD 612522; 1:1,500), Rabbit α BLM (Abcam ab2179; 1:2,000),
Techniques: Sequencing, Expressing, Mutagenesis